Introduction
XelToFab — Design field post-processing pipeline
XelToFab converts scalar design fields and callable SDF functions into clean, fabrication-ready triangle meshes and contours. It handles optional SDF evaluation plus the full post-processing pipeline: preprocessing, mesh extraction, smoothing, repair, and quality analysis.
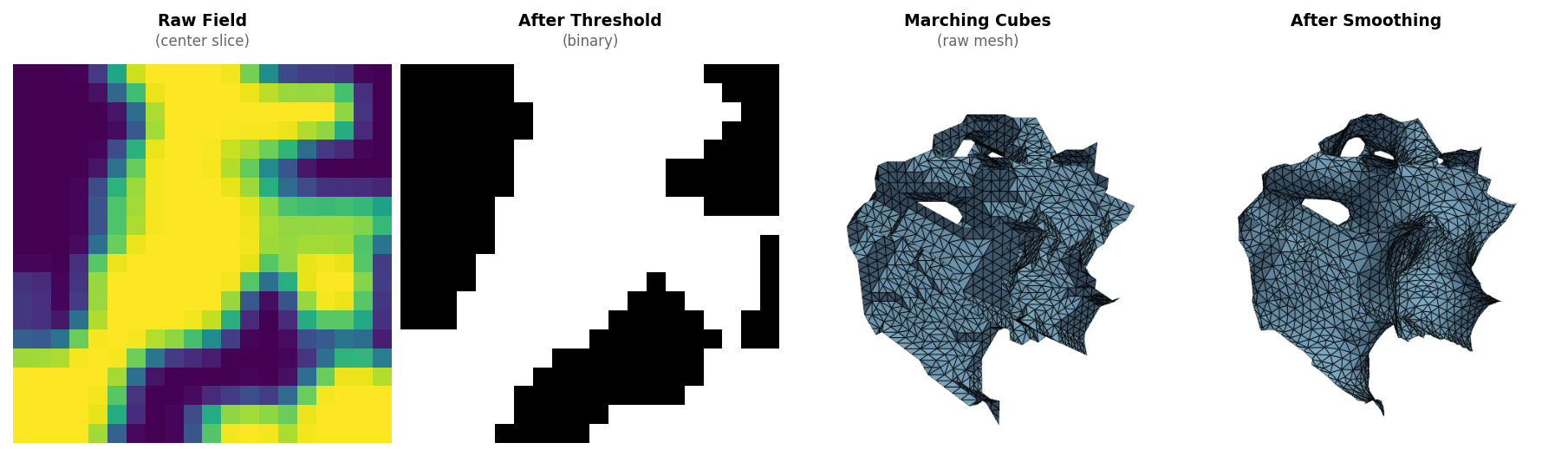
What it does
Given either a scalar field (a numpy array) or an SDF function (analytical or neural), XelToFab:
- Evaluates SDF functions (optional) — sample analytical or neural SDFs on a uniform or octree-adaptive grid via
process_from_sdf() - Preprocesses grid fields (optional) — Gaussian smoothing, thresholding, morphological cleanup
- Extracts geometry — marching cubes, dual contouring, surface nets, or manifold3d (3D → triangle mesh) or marching squares (2D → contours, or optionally an extruded 3D prism via
extrude_2d()) - Smooths the result — Taubin or bilateral filtering to remove staircase artifacts
- Repairs the mesh (optional) — non-manifold fixing for watertight geometry
- Remeshes for quality (optional) — isotropic remeshing for FEA-ready elements
- Decimates to target (optional) — quadric edge collapse to reduce face count
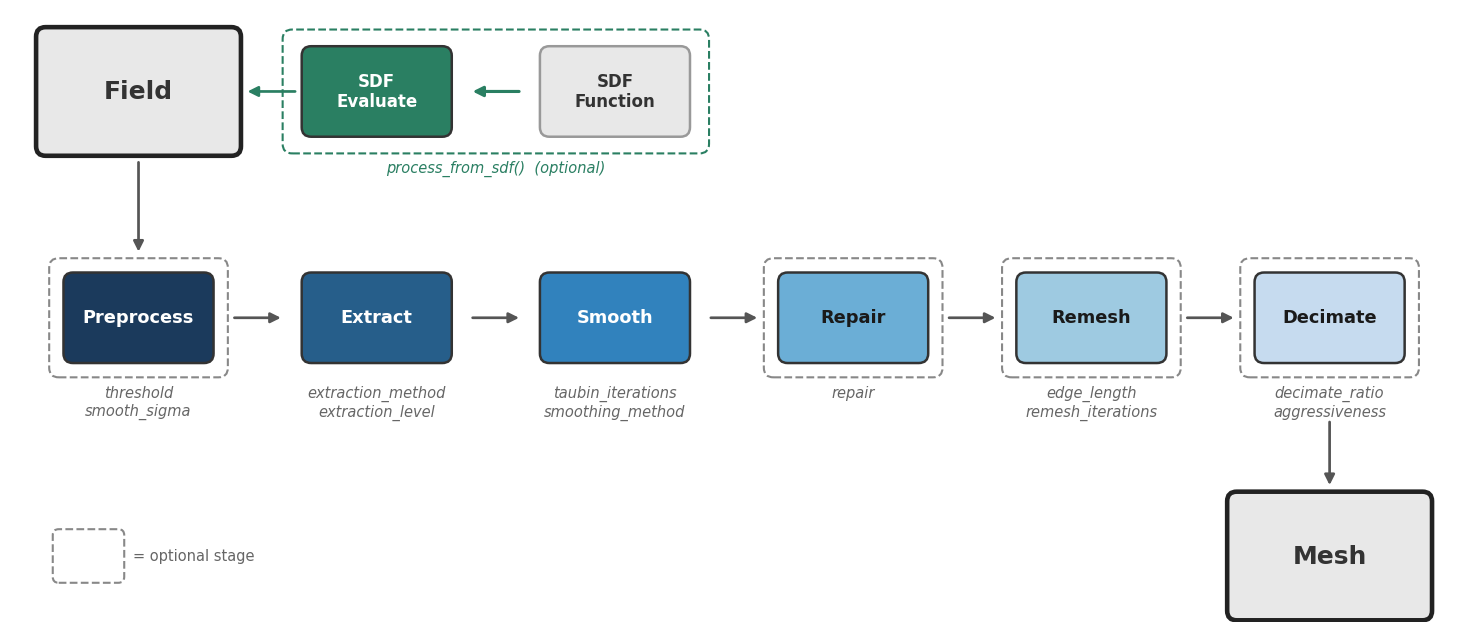
Key features
- Multi-format input — NumPy, MATLAB, VTK, HDF5, CSV
- 2D and 3D — contour extraction, 2D→3D extrusion, or native triangle mesh generation
- Multiple field types — density fields, SDFs, occupancy fields, and neural field outputs
- SDF function evaluation — process neural models and analytical SDFs directly via
process_from_sdf()with optional octree acceleration - Quality metrics — aspect ratio, min angle, scaled Jacobian via PyVista
- CLI and Python API —
xtfcommand-line tool or import directly
Quick examples
From a grid file (density or SDF array):
from xeltofab import load_field, process, save_mesh
result = process(load_field("field.npy"))
save_mesh(result, "output.stl")From an SDF function (neural model, analytical formula):
from xeltofab import process_from_sdf, save_mesh
def my_sdf(points): # [N, 3] → [N] signed distances
return np.linalg.norm(points, axis=1) - 1.0
result = process_from_sdf(my_sdf, bounds=(-2, -2, -2, 2, 2, 2), resolution=64)
save_mesh(result, "sphere.stl")Next steps
- Installation — set up XelToFab
- Quick Start — process your first field
- Pipeline Overview — understand how the pipeline works
- SDF Functions — evaluate neural and analytical SDFs